BioImpacts. 7(4):247-254.
doi: 10.15171/bi.2017.29
Original Research
Efficient and stable transformation of Dunaliella pseudosalina by 3 strains of Agrobacterium tumefaciens
Jaber Dehghani 1, 2, Ali Movafeghi 1, *, Abolfazl Barzegari 2, 3, Jaleh Barar 2, * 
Author information:
1Department of Plant Biology, Faculty of Natural Science, University of Tabriz, 29th Bahman Blvd, Tabriz, Iran
2Research Center for Pharmaceutical Nanotechnology, Tabriz University of Medical Science, Tabriz, Iran
3School of Advanced Biomedical Sciences, Tabriz University of Medical Science, Daneshgah street, Tabriz, Iran
Abstract
Introduction:
Several platforms including mammalian, plant and insect cells as well as bacteria, yeasts, and microalgae are available for the production of recombinant proteins. Low efficiency of delivery systems, extracellular and intracellular degradation of foreign genes during transformation, difficulties in targeting and importing into the nucleus, and finally problems in integration into nuclear genome are the most bottlenecks of classical plasmids for producing recombinant proteins. Owing to high growth rate, no common pathogen with humans, being utilized as humans’ food, and capability to perform N-glycosylation, microalgae are proposed as an ideal system for such biotechnological approaches. Here, Agrobacterium tumefaciens is introduced as an alternative tool for transformation of the microalga Dunaliella pseudosalina.
Methods:
The transformation of gfp gene into the D. pseudosalina was evaluated by three strains including EHA101, GV3301 and GV3850 of A. tumefaciens. The integrating and expression of gfp gene were determined by PCR, RT-PCR, Q-PCR and SDS-PAGE analyses.
Results:
The T-DNA of pCAMBIA1304 plasmid was successfully integrated into the genome of the microalgal cells. Although all of the strains were able to transform the algal cells, GV3301 possessed higher potential to transform the microalgal cells in comparison to EHA101 and GV3850 strains. Moreover, the stability of gfp gene was successfully established during a course of two months period in the microalgal genome.
Conclusion
: Agrobacterium is introduced as a competent system for stable transformation of Dunaliella strains in order to produce eukaryotic recombinant proteins.
Keywords:
grobacterium tumefaciens
, Dunaliella pseudosalina, Gene expression, Microalgal transformation
Copyright and License Information
© 2017 The Author(s)
This work is published by BioImpacts as an open access article distributed under the terms of the Creative Commons Attribution License (
http://creativecommons.org/licenses/by-nc/4.0/). Non-commercial uses of the work are permitted, provided the original work is properly cited.
Introduction
To date, five common platforms including mammalian cells, bacteria, yeasts, insect cells and plant cells have frequently been used for the production of recombinant proteins.
1
The absence of post-transcriptional and post-translational modifications in bacteria, low production efficiency in mammalians, hyper-mannosylation during N-glycan processing in yeasts, the existence of terminal mannose and α-1, 3 fucose in N-glycans of insects and the presence of xylose (not seen in human) and α-1,3 fucose in plant cell glycoproteins are the major limitations of these organisms.
1,2
Recently, microalgae are introduced as an ideal system for production of recombinant proteins. Microalgae are a wide group of photosynthetic microorganisms that fix global organic carbon around 50%.
3
In addition, they have high potentials to produce vital compounds such as pigments (β-carotene and astaxanthin,), vitamins, proteins and vital lipids (e.g. eicosapentaenoic acid and docosahexaenoic acid) as dietary supplements.
4
Furthermore, fast-growing pattern, low food requirements (i.e., only light, water and basic nutrients), photo autotrophy and ease of cultivation are a number of other advantages of the microalgae.
4,5
Importantly, eukaryotic microalgae are capable of post-translational modification such as N-glycosylation in expressed proteins almost similar to human glycoproteins.
6
In addition, many of microalgae are considered as generally safe for human (GRAS) and their recombinant proteins could be used without purification.
5
Up to now, several microalgae especially Chlamydomonas reinhardtii
7
, Phaeodactylum tricornutum,
8
Dunaliella salina
9
and Chlorella species
10
have been successfully transformed to produce recombinant proteins. Different methods such as electroporation and particle bombardment are used to deliver the heterologous genes into the microalgae genomes.
7,11
Except D. salina, the other mentioned microalgae have a very rigid cell wall, causing very low transformation competence and unstable transgene integrations.
12
In fact, Dunaliella species with fast growth rate and without cell wall seem to be an ideal platform for transformation studies.
13
Currently, different synthetic plasmids are introduced for the transformation of some microalgae.
9,14
However, owing to high potential risks of intra/extra degradation and difficulties of gene introducing process, these approaches could not be succeeded sufficiently for microalgae transformation.
15
Recently, Agrobacterium tumefaciens-mediated transformations are presented as a stable and reliable tool for a few microalgae such as C. renhardtii
16
and Haematococcus pluvialis
17
with highly efficient frequency. Here, we successfully transformed D. pseudosalina isolate MAH (canthaxanthin producing strain) for the first time. The transformation and expression efficiency of green florescence protein gene (gfp) were evaluated by using three strains of A. tumefaciens.
Material and Methods
Materials
The antibiotics (hygromycin, kanamycin, rifampicin and cefotaxime) were purchased from Sigma-Aldrich Company (Missouri, United States). Taq DNA Polymerase Master Mix RED for PCR experiments was obtained from Ampliqon Company (Odense, Denmark). Moreover, all of the reagents and enzymes that used for RNA extraction and cDNA synthesis were purchased from Invitrogen Company (California, USA). SYBR green master mix for Q-PCR is also obtained from Qiagen Company (Hilden, Germany).
Microalga isolation and medium
Dunaliella pseudosalina isolate MAH was collected from the saline Lake Maharlu (Fars province, Iran) and was morphologically and molecularly identified (Accession no: KU641615.1). For transformation procedures and selection of the transformants, the microalgae were grown in Walne’s medium.
18
Subsequently, the microalgae were incubated at 25°C, light/dark cycle (16:8) and 80 μmol photon m-2s-1 irradiance for downstream processes.
Bacterial strains
Among different strains of A. tumefaciens, which are commonly used for the plant genetic transformation, three strains were experienced for the transformation of algal cells. The used strains of A. tumefaciens were dissimilar based on their non-oncogenic plasmid structure. Tree strains of A. tumefaciens include EHA101 (Nopaline type, pTiBo542DT-DNA plasmid), GV3850 (Nopaline type, pGV3850 plasmid) and GV3301 (Nopaline type, pMP90 plasmid). The strains of A. tumefaciens were purchased from Genetics and Agricultural Biotechnology Institute of Tabarestan, Sari, Iran. All strains were transformed using pCAMBIA1304 via the freeze-thaw method and were kept on LB medium containing 100 mg/L kanamycin. The sequence of pCAMBIA 1304 plasmid was handled by SnapGene Viewer software.
Plasmid structure
The Agrobacterium-based plasmid, pCAMBIA1304, was used for transformation of algal cells as one of the most routine Cambia plasmids. This plasmid contains kanamycin resistance gene for bacterial selection, hygromycin resistance gene (hpt) for plant cell selection and gfp-uidA fusion as a reporter gene under the action of the CaMV35S promoter (Fig. 1). The pCambia1304 vector has small size (12362 bp) and high copy number in E.coli for high DNA yields. E. coli strain DH5α was used for the maintenance of the plasmid.
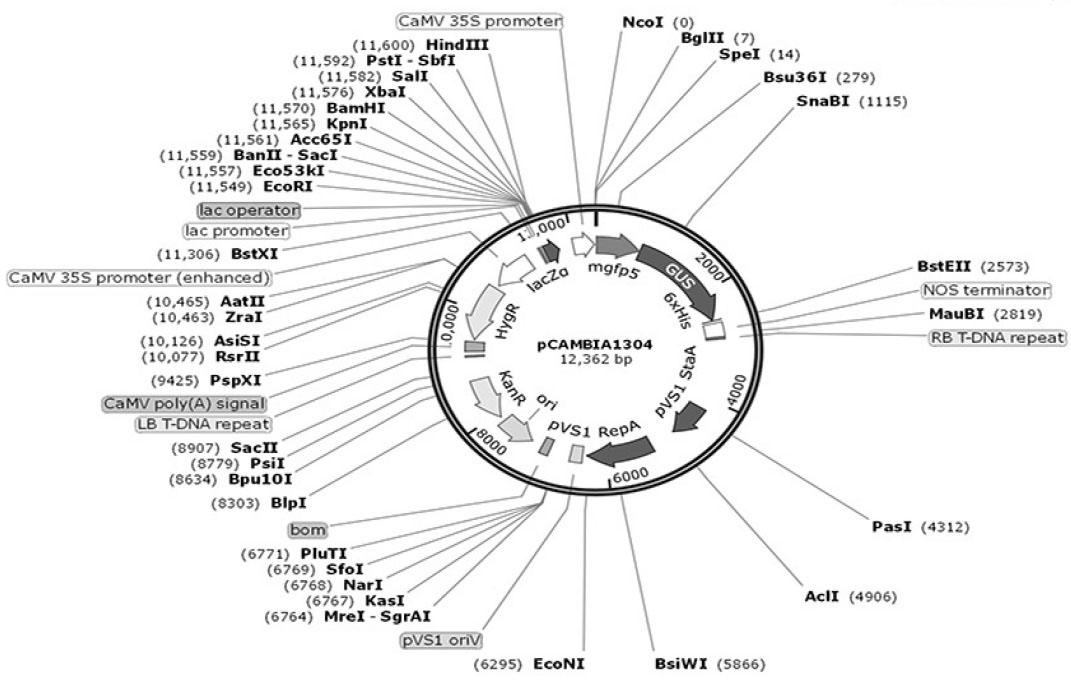
Fig. 1.
The structure of pCAMBIA1304 plasmid. T-DNA of the plasmid contains hpt and fusion gfp-gus genes that both are driven by
CaMV35S promoter.
.
The structure of pCAMBIA1304 plasmid. T-DNA of the plasmid contains hpt and fusion gfp-gus genes that both are driven by
CaMV35S promoter.
Antibiotic sensitivity test
Approximately 1 × 107 cells per ml (OD600: 0.8) of D. pseudosalina were plated and spread on solid medium containing 100, 150 and 200 mg/L hygromycin to determine the appropriate concentration of the antibiotic for the selection of transformants. Solid medium without hygromycin was used as a control. The observations of antibiotic sensitivity tests were made after 14 days. These experiments were performed in four repeats.
Microalgae transformation procedures
Dunaliella pseudosalina is transformed by co-cultivating microalgal cells with A. tumefaciens strains.
16
In brief, 150 µL of microalgal cultures in the log phase of growth were plated on the solidified Walne’s medium and incubated in light for 10 days to form the algal lawn. An overnight grown Agrobacterium strains in liquid LB medium (at 28°C) containing an appropriate combination of antibiotics (20 mg/L rifampicin and 50 mg/L kanamycin) were centrifuged at 6000 g for 5 minutes, and finally, the bacterial pellets were resuspended in liquid Walne’s medium. Subsequently, 200 µL of the bacterial suspensions were spread to the thin layer of D. pseudosalina lawn grown on agar plates. Subsequently, the co-cultivation plates were incubated at 25°C for 48 hours. Finally, microalgal cells were harvested by centrifugation at 1000g for 2 min and washed three times with liquid Walne’s medium containing 500 mg/L cefotaxime. The washed microalgal cells were cultured on the solidified Walne’s medium containing 200 mg/L hygromycin and 500 mg/L cefotaxime. After 12 days, the hygromycin-resistant colonies appeared in the selection medium. Individual colonies were picked and transferred into the liquid Walne’s medium for the following processing analyses.
DNA analysis
The microalgal cells were harvested in the late logarithmic phase of growth by centrifugation (2500 g for 10 minutes) and then genomic DNA extraction was performed using the CTAB based method.
19
The presence of hpt and gfp genes and the absence of kanamycin resistance gene (kan) in D. pseudosalina genome were confirmed by polymerase chain reaction (PCR) using specific primers (Table 1), designed by Oligo software (version 7). The oligonucleotide primer sequences were purchased from Takapo Zist Company, Tehran, Iran. PCR thermocycling condition was as follows: initial denaturation at 94˚C for 5 min, annealing at 52-57˚C for 1 minute and extension at 72˚C for 1 minute, for 32 successive cycles followed by a final extension step at 72°C for 5 minutes. The PCR products were analyzed using 1% agarose gel electrophoresis.
RNA extraction and RT-PCR
Total RNA content from transformed and untransformed (as the negative control) D. pseudosalina was extracted using TRIzol® reagent according to the instruction of the manufacturer. The purity of RNA was assessed by evaluation the A260/280 ratio using NanoDrop ND-1000 spectrophotometer (Thermo Fisher Scientific, U.S.A.). For cDNA synthesis, 2 μg of total RNAs, 1 μL random hexamer primer, 1 μL dNTP and UltraPureTM DEPC-Treated water were mixed and incubated at 65 °C for 5 minutes and then transferred onto the ice. Then, MMLV reverse transcriptase (5 U), 1x reverse transcription (RT) buffer, and 1 U/L RNase inhibitor were added to the reaction mixture and UltraPureTM DEPC-Treated Water was used to obtain a final volume of 20 μL. Further, RT was performed for 10 minutes at 25°C followed by 1 h at 42°C. Real-time PCR reactions with q-gfp specific primer (Table 1) were performed to measure the expression level of gfp gene using SYBRgreen master mix.
Table 1.
The list of used primers
|
Primers
|
Forward sequence
|
Reverse sequence
|
T
m
(°C)
|
|
gfp
|
ATGGTAGATCTGACTAGTAAAGG |
TCATCCATGCCATGTGTAATC |
52 |
|
hpt
|
ATGAAAAAGCCTGAACTCACCG |
CTATTTCTTTGCCCTCGGACG |
56 |
|
kan
|
ATGGCTAAAATGAGAATATCACCGG |
GGCTTGATCCCCAGTAAGTC |
56 |
|
q-gfp
|
GCACAAATTTTCTGTCAGTGGAG |
CGTGCCGCTTCATATGATCTG |
57 |
Abbreviation: Tm, melting temperature.
SDS-PAGE experiments
Total soluble proteins were extracted from transformed and untransformed microalgal cells by extraction buffer (50 mM Na2HPO4, pH 7.0, 10 mM Na2EDTA, 2 mM β-mercaptoethanol, 2 mM phenylmethylsulfonyl fluoride (PMSF)) and sonicated (12 kHz) on ice for 10 minutes.
16
After sonication, the lysates were centrifuged at 12000 g for 5 minutes and the supernatants were transferred to new tubes. The total protein concentration was determined by absorbance at 280 nm using NanoDrop ND-1000 spectrophotometer. Finally, equal amounts (20 mg) of total soluble protein were subjected to SDS-PAGE using 10% gels. The visualizing of proteins on the gels was performed with silver nitrate staining procedure.
20
Results
Transformation efficiency of Agrobacterium strains
Our antibiotic sensitivity experiments revealed that 200 mg/L hygromycin could completely inhibit the growth of D. pseudosalina on the solid plates (Fig. 2). It is established that the GV3301, EHA101 and GV3850 strains of Agrobacterium have respectively high medium requirements and very low potential for transformation (Fig. 3). Moreover, the best concentration of Agrobacterium in transformation process is determined as OD600: 0.7. However, the higher concentration of Agrobacterium killed the microalgal colonies.
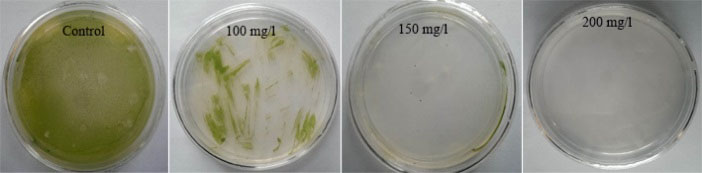
Fig. 2.
Hygromycin sensitivity tests. Three concentrations of hygromycin are used for determining the suitable antibiotic concentration to select transformed microalgal colonies. 200 mg/L of hygromycin could completely inhibit the growth of D. pseudosalina colonies.
.
Hygromycin sensitivity tests. Three concentrations of hygromycin are used for determining the suitable antibiotic concentration to select transformed microalgal colonies. 200 mg/L of hygromycin could completely inhibit the growth of D. pseudosalina colonies.
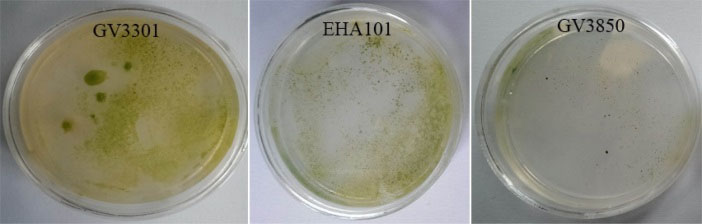
Fig. 3.
Transformation efficacy rate via Agrobacterium strains. GV3301 strain owns the highest and GV3850 has the lowest transformation efficiency.
.
Transformation efficacy rate via Agrobacterium strains. GV3301 strain owns the highest and GV3850 has the lowest transformation efficiency.
Integration of T-DNA into the genome of D. pseudosalina
Five days after transformation, PCR with genomic DNA and the specific primers of hpt and gfp genes have respectively given expected amplified bands of ~1000 and ~700 bp in the D. pseudosalina colonies that were transformed by the Agrobacterium strains. In comparison, the control untransformed colonies resulted in no band (Fig. 4). Additionally, PCR-based evaluation showed the stable presence of gfp gene in D. pseudosalina genome during 6 weeks after transformations (Fig. 5).
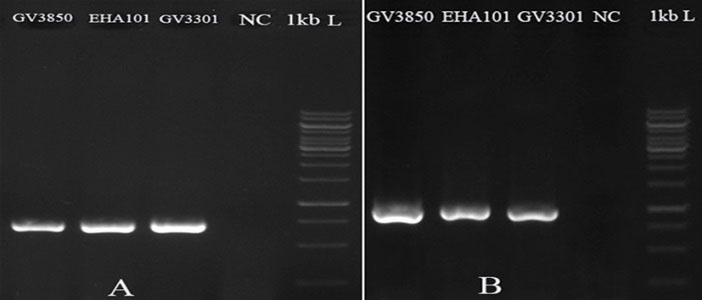
Fig. 4.
Integration of T-DNA into the transformed D. pseudosalina cells. gfp gene has 700 bp (A) and hpt gene has 1000 bp size (B). Untransformed microalgal cells are used as negative control (NC).
.
Integration of T-DNA into the transformed D. pseudosalina cells. gfp gene has 700 bp (A) and hpt gene has 1000 bp size (B). Untransformed microalgal cells are used as negative control (NC).
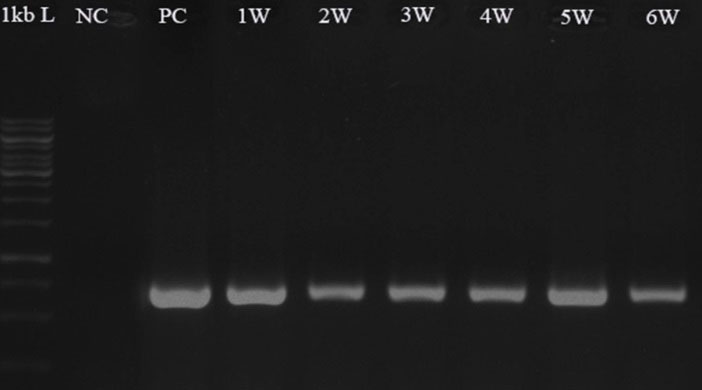
Fig. 5.
Confirming the presence of gfp gene in the transformed D. pseudosalina colonies during 6 weeks after transformation. Untransformed microalga is used as negative control (NC). Purified pCAMBIA1304 plasmid is used as positive control (PC).
.
Confirming the presence of gfp gene in the transformed D. pseudosalina colonies during 6 weeks after transformation. Untransformed microalga is used as negative control (NC). Purified pCAMBIA1304 plasmid is used as positive control (PC).
Confirmation of gfp gene expression
To determine the expression of gfp gene, RT-PCR reaction was performed on transformed D. pseudosalina. The electrophoresis showed a band of ~700 bp in all transformed microalgal colonies by Agrobacterium strains. Genomic DNA of transformed colonies was used as a positive control (Fig. 6A). Moreover, the expression of gfp gene in the transformed D. pseudosalina colonies by GV3301 of Agrobacterium was proved with specific primer by Q-PCR (Fig. 6B).
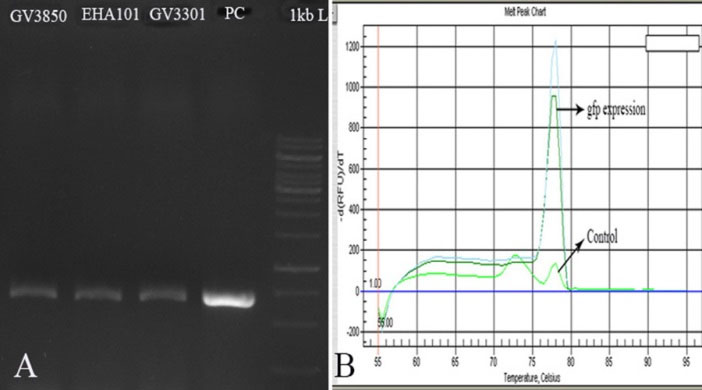
Fig. 6.
Confirming the expression of gfp gene by RT-PCR (A) and Q-PCR (B). In RT-PCR, genomic DNA of transformed D. pseudosalina is used as positive control. In Q-PCR, cDNA of untransformed microalgal cells are used as control.
.
Confirming the expression of gfp gene by RT-PCR (A) and Q-PCR (B). In RT-PCR, genomic DNA of transformed D. pseudosalina is used as positive control. In Q-PCR, cDNA of untransformed microalgal cells are used as control.
The expression of the protein product from the introduced gfp gene in the transformed D. pseudosalina colonies was further evaluated by SDS-PAGE analysis. The results revealed a protein band with a molecular weight of about 90 kDa in the transformants (Fig. 7). Regarding the pCAMBIA1304 plasmid, the ~90 kDa band presents the GFP-GUS fusion protein.
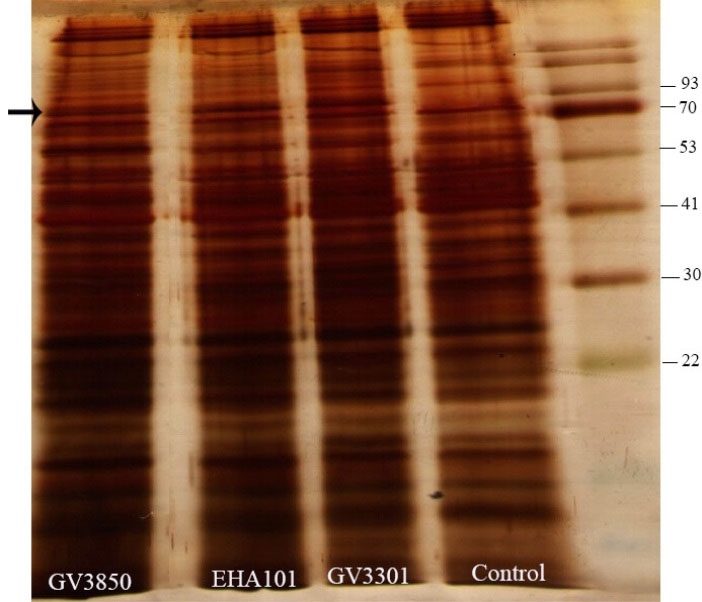

Fig. 7.
Detection of the product of gfp-gus fusion gene by SDS-PAGE. A ~90 KD protein band is existed in transformed D. pseudosalina cells. This band is absent in untransformed microlagal cells.
.
Detection of the product of gfp-gus fusion gene by SDS-PAGE. A ~90 KD protein band is existed in transformed D. pseudosalina cells. This band is absent in untransformed microlagal cells.
To determine the probable Agrobacterium contaminations, PCR reactions were performed using the specific primer of kan gene (located outside of T-DNA in pCAMBIA1304) and the genomic DNA of transformed D. pseudosalina colonies. The results revealed no amplification in the transformed colonies with primers of kan gene, while PCR with pCAMBIA1304 as a positive control and the primer revealed a ~750 bp band in the electrophoresis gel (Fig. 8).
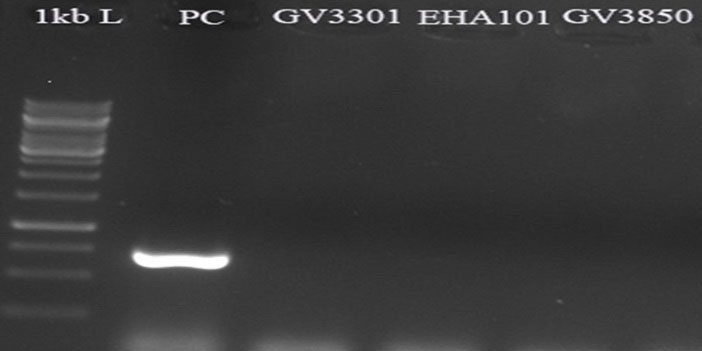
Fig. 8.
Approving no Agrobacterium contaminations by kan gene specific primer. pCAMBIA1304 plasmid is used as positive control. All of the transformed D. pseudosalina colonies show no Agrobacterium contaminations.
.
Approving no Agrobacterium contaminations by kan gene specific primer. pCAMBIA1304 plasmid is used as positive control. All of the transformed D. pseudosalina colonies show no Agrobacterium contaminations.
Discussion
Nowadays, different methods have been used for the transformation of microalgae. High-voltage electrical pulses for transiently permeabilization cell wall and cell membranes, known as electroporation, is routinely used to deliver plasmid or DNA to different organisms, including mammalian,
21
yeast,
22
bacteria,
23
plant protoplasts
24
and different species of microalgae
25
as well. Although, electroporation is an efficient tool for gene transfer, high mortality of cells after exposure to high-voltage as well as difficulties for in vivo application, are considered as the major shortcomings of this method.
15
Agitating the mixture of plasmid DNA and glass beads in the presence of polyethylene glycol (PEG) is another approach for transformation of microalgae.
26,27
PEG generally has a positive charge and binds to the plasmid. Moreover, the organisms have different types of negative charges, so that PEG allows the plasmid and organisms to get closer together. But because of high fatality rate of cells and the presence of rigid cell wall, the efficiency of this method is very low in many of microalgae.
Particle bombardment is also regarded as a capable technique to introduce the DNA into the cell walled organisms such as plants and microalgae. In this process, DNA is coated with gold or tungsten microparticles to accelerate the velocity of penetration into cell membranes or cell walls. However, this procedure is expensive, while the entry pathway of DNA and the location of DNA insertion are out of the control.
15
Another main issue is the concerns about degradation of plasmid or DNA before introducing into cells. Although the extracellular degradation can be inhibited by complex formation with various polymers and lipids,
15
this approach is not yet employed in microalgae transformations. On the other hand, after delivering the plasmids into the cytosol of target cells, it is essential to protect them from degradation by nucleases. Overall, cytosol is an unfavorable environment for plasmid DNA.
15
Some plasmids are designed to operate without interaction with native cell proteins. Under these conditions, the risk of plasmid degradation by the organism defense systems is very high.
28,29
Moreover, performing the nuclear-independent transcription and translation processes generally result in the production of low-quality proteins.
Movement the plasmids (or DNAs) in the cytosol and finding the nucleus are the final processes for DNA delivery. Plasmid movement through the cytosol toward the nucleus is probably performed by diffusion which is relatively slow.
15
For targeting and importing DNA molecules into the nucleus, the presence of nuclear localization signals (NLSs) is needed on the DNA.
30
However, this tool is not still used in microalgae transformation experiments.
Because of all difficulties, the development of alternative methods with high transformation and expression efficacy is extremely required. Recently, A. tumefaciens based plasmids are investigated to transform a number of organisms including plant, yeast, fungi and microalgae.
31
A. tumefaciens is a gram-negative soil bacterium causing the crown gall disease in plants.
32
Infection process by this bacterium is mediated by the presence of a large size plasmid (200-800 kb) known as pTi.
31
This plasmid contains one or more DNA fragments called transfer-DNA (T-DNA), which enter into the plant cells and cause the disease.
31
After importing the T-DNA into the plant cell, the products of virD2 and virE2 genes (located on the pTi plasmid) are respectively connected to 5' end and covered the whole length of T-DNA. These proteins inhibit degradation of T-DNA by the plant cell endonucleases. Beside, VirD2 and VirE2 contain NLS sequences allowing T-DNA to be directed to the nucleus. In addition, it is believed that VirE3 helps nuclear localization of VirE2.
31
However, it is known that some of the host genes are required for this process to be accomplished.
32
In Arabidopsis, for example, VirD2 protein interacts with AtKAPα (as known importin α) and VirE2 interacts with the plant VirE2 interacting protein 1 (VIP1) and also VirE3 protein. VIP1 and VirE3 act as molecular adaptors between VirE2 and importin α, enabling VirE2 to be transferred into the nucleus.
33
Finally, VirF protein of Agrobacterium mediates the uncoating process of T-DNA in the nucleus.
34
Integration of the T-DNA into the plant genome occurs by illegitimate recombination into the transcriptionally active regions,
32
but the precise mechanism of this process is still unclear. Although, it seems that VirD2, VIP1 and H2A proteins of plant cells are effective to integrate the T-DNA into the nuclar genome.
35
Furthermore, Agrobacterium based plasmids with high transformation rate, enable the gene of interest to escape from host cell endonucleases and also allow the exclusive nuclear integration of the genes, hence could be considered as a potentially powerful alternative platform for the transformation of different microalgae.
36
Additionally, regarding the size of T-DNA (30-50 kb),
32
Agrobacterium-based plasmids allow loading/carrying the high size genes.
In the present work, we used EHA101, GV3301 and GV3850 strains of A. tumefaciens, carrying pCAMBIA1304 plasmid, to transform the microalga D. pseudosalina. Our PCR experiments confirmed the successful integration of gfp and hpt genes into the D. pseudosalina genome. The expression of gfp gene was verified by RT-PCR and Q-PCR and consequently the band of GFP-GUS fusion protein was visualized by SDS-PAGE analysis. The stability of gfp gene in the transformants was confirmed during a course of two months period after transformation. Among EHA101, GV3301 and GV3850 strains of A. tumefaciens, only EHA101 strain is previously used for transformation of a few microalgae. For instance, Haematococcus pluvialis was transformed by EHA101 strain of A. tumefaciens and the expression of gfp, gus and hpt genes were confirmed. Besides, different microscopical and molecular analyses revealed the stable integration of the genes in the nuclear genome of H. pluvialis followed by the expression of integrated gene product.
17
Additionally, the mentioned strain is employed for transformation of Dunaliella bardawil and expression of gfp gene.
37
Although the GV3301 and GV3850 strains of A. tumefaciens were frequently used for transformation of different higher plants such as Zea mays, Brassica juncea and Vicia narbonensis.
38-40
these strains have not been applied for microalgae transfection. In bean (Phaseolus vulgaris L.), for example, the GV3301 strain show higher potential than EHA105 (a kanamycin sensitive type of EHA101) regarding the transformation and expression efficiency.
41
Similar findings were reported in Arabidopsis.
42
Intriguingly, our data showed that the mentioned strains could be successfully employed for stable transformation of D. pseudosalina. We showed that GV3301 strain possesses higher potential to transform the microalgal cells than EHA101 and GV3850 strains. Probably, the GV3301 strain can strongly bind to the D. pseudosalina cells causing high transformation efficacy. However, all of the Agrobacterium strains seem to be competent to integrate the genes of interest. Accordingly, other strains of A. tumefaciens were used to transform some microalgae species in order to evaluate this approach for producing recombinant proteins. For instance, the LBA4404 strain of A. tumefaciens is used to transform C. reinhardtii by pCAMBIA1304. Molecular, histochemical and microscopical analysis confirmed the stable integration and the expression of T-DNA in the transformed colonies. Moreover, the transformation frequency efficiency of this method was 50 fold higher than the glass beads based transfection.
16
Later, the introducing of pCAMBIA1301 into C. reinhardtii, revealed that EHA105 strain of A. tumefaciens is more competent than LBA4404.
43
Transformation and expression of egfp and gus genes in Schizochytrium protoplasts, a docosahexaenoic acid producing marine microalgae, were also studied by A. tumefaciens.
44
It was reported that LBA4404 strain of A. tumefaciens processes higher potential to transform the microalgae than the EHA105 strain.
44
Conclusion
In this research, we set up a successful procedure in order to stable and efficient transformation of the microalga D. pseudosalina by three A. tumefaciens strains. Dunaliella species have high salinity (0.5-5 M NaCl) habitat that enables the algal growth in open ponds without any concerns about dangerous contaminations. Since microalgae can produce high quality recombinant proteins, handling of A. tumefaciens strains for production of vital recombinant proteins in Dunaliella strains is highly suggested.
Ethical approval
There is none to be declared.
Competing interests
There is no conflict of interests to be reported.
Acknowledgments
Authors are grateful for the financial support provided by Research Center for Pharmaceutical Nanotechnology at Tabriz University of Medical Sciences.
Research Highlights
What is current knowledge?
simple
-
√ Agrobacterium tumefaciens strains especially GV3301
are more powerful approach for stable and efficient
transformation of Dunaliella pseudosalina.
What is new here?
simple
-
√ This is the first report about transformation of the microalga
Dunaliella pseudosalina.
-
√ The current work is the first successful study about
applying GV3301 and GV3850 strains of A. tumefaciens for
stable transformation of microalgae.
References
- Dingermann T. Recombinant therapeutic proteins:production platforms and challenges. Biotechnol J 2008; 3:90-7. doi: 10.1002/biot.200700214 [Crossref] [ Google Scholar]
- Graumann K, Premstaller A. Manufacturing of recombinant therapeutic proteins in microbial systems. Biotechnol J 2006; 1:164-86. doi: 10.1002/biot.200500051 [Crossref] [ Google Scholar]
- Field CB, Behrenfeld MJ, Randerson JT, Falkowski P. Primary production of the biosphere:integrating terrestrial and oceanic components. Science 1998; 281:237-40. doi: 10.1126/science.281.5374.237 [Crossref] [ Google Scholar]
- Leon-Banares R, Gonzalez-Ballester D, Galvan A, Fernandez E. Transgenic microalgae as green cell-factories. Trends Biotechnol 2004; 22:45-52. doi: 10.1016/j.tibtech.2003.11.003 [Crossref] [ Google Scholar]
- Potvin G, Zhang Z. Strategies for high-level recombinant protein expression in transgenic microalgae:a review. Biotechnol Adv 2010; 28:910-8. doi: 10.1016/j.biotechadv.2010.08.006 [Crossref] [ Google Scholar]
- Mathieu-Rivet E, Kiefer-Meyer MC, Vanier G, Ovide C, Burel C, Lerouge P. Protein N-glycosylation in eukaryotic microalgae and its impact on the production of nuclear expressed biopharmaceuticals. Front Plant Sci 2014; 5:359. doi: 10.3389/fpls.2014.00359 [Crossref] [ Google Scholar]
- Rasala BA, Lee PA, Shen Z, Briggs SP, Mendez M, Mayfield SP. Robust expression and secretion of Xylanase1 in Chlamydomonas reinhardtii by fusion to a selection gene and processing with the FMDV 2A peptide. PLoS One 2012; 7:e43349. doi: 10.1371/journal.pone.0043349 [Crossref] [ Google Scholar]
- Hempel F, Lau J, Klingl A, Maier UG. Algae as protein factories:expression of a human antibody and the respective antigen in the diatom Phaeodactylum tricornutum. PLoS One 2011; 6:e28424. doi: 10.1371/journal.pone.0028424 [Crossref] [ Google Scholar]
- Geng D, Wang Y, Wang P, Li W, Sun Y. Stable expression of hepatitis B surface antigen gene in Dunaliella salina (Chlorophyta). J Appl Phycol 2003; 15:451-6. doi: 10.1023/B:JAPH.0000004298.89183.e5 [Crossref] [ Google Scholar]
- Koo J, Park D, Kim H. Expression of bovine lactoferrin N-lobe by the green alga, Chlorella vulgaris. Algae 2013; 28:379-87. doi: 10.4490/algae.2013.28.4.379 [Crossref] [ Google Scholar]
- Hempel F, Maier UG. An engineered diatom acting like a plasma cell secreting human IgG antibodies with high efficiency. Microb Cell Fact 2012; 11:1. doi: 10.1186/1475-2859-11-126 [Crossref] [ Google Scholar]
- Cadoret JP, Bardor M, Lerouge P, Cabigliera M, Henriquez V, Carlier A. Microalgae as cell factories producing recombinant commercial proteins. Med Sci (Paris) 2008; 24:375-82. doi: 10.1051/medsci/2008244375 [Crossref] [ Google Scholar]
- Barzegari A, Hejazi MA, Hosseinzadeh N, Eslami S, Mehdizadeh Aghdam E, Hejazi MS. Dunaliella as an attractive candidate for molecular farming. Mol Biol Rep 2010; 37:3427-30. doi: 10.1007/s11033-009-9933-4 [Crossref] [ Google Scholar]
- Pollock SV, Mukherjee B, Bajsa-Hirschel J, Machingura MC, Mukherjee A, Grossman AR. A robust protocol for efficient generation, and genomic characterization of insertional mutants of Chlamydomonas reinhardtii. Plant Methods 2017; 13:22. doi: 10.1186/s13007-017-0170-x [Crossref] [ Google Scholar]
- Luo D, Saltzman WM. Synthetic DNA delivery systems. Nature biotechnol 2000; 18:33-7. doi: 10.1038/71889 [Crossref] [ Google Scholar]
- Rajam MV, Kumar SV. Green Alga (Chlamydomonas reinhardtii). Methods Mol Biol 2006; 344:421-33. doi: 10.1385/1-59745-131-2:421 [Crossref] [ Google Scholar]
- Kathiresan S, Chandrashekar A, Ravishankar GA, Sarada R. Agrobacterium-mediated transformation in the green alga Haematococcus pluvialis (chlorophyceae, volvocales). J Phycol 2009; 45:642-9. doi: 10.1111/j.1529-8817.2009.00688.x [Crossref] [ Google Scholar]
- Raja R, Hemaiswarya S, Balasubramanyam D, Rengasamy R. Protective effect of Dunaliella salina (Volvocales, Chlorophyta) against experimentally induced fibrosarcoma on wistar rats. Microbiol Res 2007; 162:177-84. doi: 10.1016/j.micres.2006.03.009 [Crossref] [ Google Scholar]
- Hejazi MA, Barzegari A, Gharajeh NH, Hejazi MS. Introduction of a novel 18S rDNA gene arrangement along with distinct ITS region in the saline water microalga Dunaliella. Saline Syst 2010; 6:4. doi: 10.1186/1746-1448-6-4 [Crossref] [ Google Scholar]
- Heukeshoven J, Dernick R. Simplified method for silver staining of proteins in polyacrylamide gels and the mechanism of silver staining. Electrophoresis 1985; 6:103-12. doi: 10.1002/elps.1150060302 [Crossref] [ Google Scholar]
- Longo AL, Kavran JM, Kim MS, Leahy DJ. Generating mammalian stable cell lines by electroporation. Methods Enzymol 2013; 529:209-26. doi: 10.1016/b978-0-12-418687-3.00017-3 [Crossref] [ Google Scholar]
- Benatuil L, Perez JM, Belk J, Hsieh CM. An improved yeast transformation method for the generation of very large human antibody libraries. Protein Eng Des Sel 2010; 23:155-9. doi: 10.1093/protein/gzq002 [Crossref] [ Google Scholar]
- Calvin NM, Hanawalt PC. High-efficiency transformation of bacterial cells by electroporation. J Bacteriol 1988; 170:2796-801. [ Google Scholar]
- Zhang HM, Yang H, Rech EL, Golds TJ, Davis AS, Mulligan BJ. Transgenic rice plants produced by electroporation-mediated plasmid uptake into protoplasts. Plant Cell Rep 1988; 7:379-84. doi: 10.1007/bf00269517 [Crossref] [ Google Scholar]
- Sun Y, Yang Z, Gao X, Li Q, Zhang Q, Xu Z. Expression of foreign genes in Dunaliella by electroporation. Mol Biotechnol 2005; 30:185-92. doi: 10.1385/mb:30:3:185 [Crossref] [ Google Scholar]
- Kindle KL. High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 1990; 87:1228-32. doi: 10.1073/pnas.87.3.1228 [Crossref] [ Google Scholar]
- Feng S, Xue L, Liu H, Lu P. Improvement of efficiency of genetic transformation for Dunaliella salina by glass beads method. Mol Biol Rep 2009; 36:1433-9. doi: 10.1007/s11033-008-9333-1 [Crossref] [ Google Scholar]
- Obbard DJ, Gordon KH, Buck AH, Jiggins FM. The evolution of RNAi as a defence against viruses and transposable elements. Philos Trans R Soc Lond B Biol Sci 2009; 364:99-115. doi: 10.1098/rstb.2008.0168 [Crossref] [ Google Scholar]
- Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P. Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol 2011; 9:467-77. doi: 10.1038/nrmicro2577 [Crossref] [ Google Scholar]
- Branden LJ, Mohamed AJ, Smith CI. A peptide nucleic acid-nuclear localization signal fusion that mediates nuclear transport of DNA. Nat Biotechnol 1999; 17:784-7. doi: 10.1038/11726 [Crossref] [ Google Scholar]
- Gelvin SB. Agrobacterium-mediated plant transformation:the biology behind the "gene-jockeying" tool. Microbiol Mol Biol Rev 2003; 67:16-37. doi: 10.1128/MMBR.67.1.16-37.2003 [Crossref] [ Google Scholar]
- McCullen CA, Binns AN. Agrobacterium tumefaciens and plant cell interactions and activities required for interkingdom macromolecular transfer. Annu Rev Cell Dev Biol 2006; 22:101-27. doi: 10.1146/annurev.cellbio.22.011105.102022 [Crossref] [ Google Scholar]
- Gelvin SB. Plant proteins involved in Agrobacterium-mediated genetic transformation. Annu Rev Phytopathol 2010; 48:45-68. doi: 10.1146/annurev-phyto-080508-081852 [Crossref] [ Google Scholar]
- Abu-Arish A, Frenkiel-Krispin D, Fricke T, Tzfira T, Citovsky V, Wolf SG. Three-dimensional reconstruction of Agrobacterium VirE2 protein with single-stranded DNA. J Biol Chem 2004; 279:25359-63. doi: 10.1074/jbc.M401804200 [Crossref] [ Google Scholar]
- Magori S, Citovsky V. Epigenetic control of Agrobacterium T-DNA integration. Biochim Biophys Acta 2011; 1809:388-94. doi: 10.1016/j.bbagrm.2011.01.007 [Crossref] [ Google Scholar]
-
Komari T, Ishida Y, Hiei Y. Plant Transformation Technology:Agrobacterium‐Mediated Transformation. Handbook of plant biotechnology 2004.
- Anila N, Chandrashekar A, Ravishankar G, Sarada R. Establishment of Agrobacterium tumefaciens-mediated genetic transformation in Dunaliella bardawil. Eur J Phycology 2011; 46:36-44. doi: 10.1080/09670262.2010.550386 [Crossref] [ Google Scholar]
- Wang Y, Fu S, Wen Y, Zhang Z, Xia Y, Liu Y. Selection of maize inbred lines with high regeneration and susceptibility to Agrobacterium tumifacience. J Genet Genomics 2007; 34:749-55. doi: 10.1016/s1673-8527(07)60084-8 [Crossref] [ Google Scholar]
- Pickardt T, Meixner M, Schade V, Schieder O. Transformation of Vicia narbonensis via Agrobacterium-mediated gene transfer. Plant Cell Rep 1991; 9:535-8. doi: 10.1007/bf00232326 [Crossref] [ Google Scholar]
- Wang Y, Fu S, Wen Y, Zhang Z, Xia Y, Liu Y. Selection of maize inbred lines with high regeneration and susceptibility to Agrobacterium tumifacien. J Genet Genomics 2007; 34:749-55. doi: 10.1016/s1673-8527(07)60084-8 [Crossref] [ Google Scholar]
- Nguyen T, Sticklen M. Genetic transformation of common bean (Phaseolus vulgaris L) with the gus color marker, the bar herbicide resistance, and the barley (Hordeum vulgare) HVA1 drought tolerance genes. Int J Agron 2012:2012. doi: 10.1155/2012/198960 [Crossref]
- Kim MJ, Baek K, Park CM. Optimization of conditions for transient Agrobacterium-mediated gene expression assays in Arabidopsis. Plant Cell Rep 2009; 28:1159-67. doi: 10.1007/s00299-009-0717-z [Crossref] [ Google Scholar]
- Pratheesh PT, Vineetha M, Kurup GM. An Efficient Protocol for the Agrobacterium-mediated Genetic Transformation of Microalga Chlamydomonas reinhardtii. Mol Biotechnol 2014; 56:507-15. doi: 10.1007/s12033-013-9720-2 [Crossref] [ Google Scholar]
- Cheng R, Ma R, Li K, Rong H, Lin X, Wang Z. Agrobacterium tumefaciens mediated transformation of marine microalgae Schizochytrium. Microbiol Res 2012; 167:179-86. doi: 10.1016/j.micres.2011.05.003 [Crossref] [ Google Scholar]