Bioimpacts. 2025;15:30032.
doi: 10.34172/bi.30032
Original Article
Effect of microRNA-141-3p, E2F3, CDK3, and KAT2B overexpression on histologic tumor grade and metastasis status in untreated breast cancer tissues
Sepideh Ebrahimian Vargahan Data curation, Formal analysis, Investigation, Validation, Visualization, Writing – original draft, 1 
Mahmood Barati Formal analysis, Investigation, Validation, 1 
Masoud Roudbari Formal analysis, 2 
Maryam Eini Investigation, Writing – review & editing, 1 
Arshad Hosseini Conceptualization, Funding acquisition, Methodology, Project administration, Resources, Supervision, Validation, Visualization, Writing – review & editing, 1, * 
Author information:
1Department of Medical Biotechnology, School of Allied Medical Sciences, Iran University of Medical Sciences, Tehran, Iran
2Biostatistics Department, School of Public Health, Iran University of Medical Sciences, Tehran, Iran
Abstract
Introduction:
Increasing evidence has reported gene expression alterations in breast cancer (BC) tissues, necessitating their investigating to highlight the molecular basis of the disease development or progression. This study investigated the expression of miR-141, E2F3, CDK3, TP53, and KAT2B, and their association with histologic grade and metastasis in BC tissues.
Methods:
The RNA expression level of miR-141, E2F3, CDK3, TP53, and KAT2B genes was analyzed in 23 BC and 23 normal tissue samples by RT-qPCR. The associations of the expression level of these genes with clinicopathological features of the BC tissue samples were evaluated. The study also explored the correlation between RNA levels of genes and miR-141.
Results:
Expression of miR-141, E2F3, CDK3, and KAT2B demonstrated significantly higher levels in BC tumor than normal tissues. TP53 expression showed an increase in tumor compared to normal tissues, although it was insignificant. Moreover, increased RNA expression of miR-141, E2F3, CDK3, and KAT2B corresponded to the advanced stage and regional metastasis of BC. Additionally, the results demonstrated a significant correlation between RNA expression levels of miR-141 with CDK3 and E2F3 with KAT2B.
Conclusion:
Our findings highlighted clinicopathologic indicators that were relevant to aggressive BC. Besides, Correlations between overexpression of miR-141, E2F3, CDK3, and KAT2B in BC tissues suggest regulatory effects. Taken together, it seems results of this study could provide evidence that dysregulation of gene expression contributes significantly to unveiling the underlying molecular basis of BC.
Keywords: Breast cancer, microRNA-141, E2F3, CDK3, KAT2B, Metastasis
Copyright and License Information
© 2025 The Author(s).
This work is published by BioImpacts as an open access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (
http://creativecommons.org/licenses/by-nc/4.0/). Non-commercial uses of the work are permitted, provided the original work is properly cited.
Funding Statement
This study was supported by Grant No (18320) from Iran University of Medical Sciences (IUMS), Tehran, Iran.
Introduction
Breast cancer (BC) is a heterogeneous and complicated disease1-3 and, the second cause of mortality by cancer in females worldwide.4,5 Based on WHO reports, the worldwide BC patients will step-up from 2 069 792 to 2 778 850 instances from 2018 to 2040, indicating the importance of further research about the disease's molecular basis.6 The current trend emphasizes the necessity for deeper insights into the molecular biology of BC.7
MicroRNAs(miRNA/miR), one of the noncoding single-stranded RNAs with a length of 18-22 nucleotides, are involved in diverse biological activities.8-10 Depending on the type of cancer, microRNAs may function as tumor suppressors or oncogenes.11-13 MiR-141, a miR-200 family member,9,14 is expressed in many human malignancies,13,15-18 including BC.19 Upregulation or downregulation of these microRNAs, as an epigenetic alteration, can regulate different cancer steps such as invasion, metastasis, relapse, and drug resistance.16 Circulating miR-141 and miR-200c are potential biomarkers of early tumor metastasis sensing in BC cells.20 However, further research is required to investigate the expression of miR-141 and its association with metastasis and histologic grade of BC patients’ tissue samples.
Several reports demonstrated the alteration in the expression of some genes during cancer development or metastasis.21-24 Identification of differentially expressed genes could shed more light on understanding the molecular basis of cancer.25 Aberrancy of the cell cycle process is one of the well-known mechanisms of oncogenesis,26 which can be activated by various mitogens, cytokines, hormones, growth factors, etc.27 yet its potential role in BC has to become more clear. E2F transcription factor 3 (E2F3), an important gene in the cell cycle placed on chromosome 6 (NC_000006.12), is a member of the E2F family28,29 and regulates the transition of G1 to S-phase in the cell cycle.30 E2F3 plays a crucial function in the progression of different kinds of cancer.31 Overexpression of E2F3 in BC can lead to tumorigenesis by centrosome amplification and dysregulated mitosis.32 Cyclin-dependent kinase 3 (CDK3), belonging to the CDKs group with serine/threonine kinase activity, is another gene that contributes significantly to the ordinance of cell cycle advancement.33,34 It has been found that up-regulation of CDK3 suppresses the migration and invasion of BC cells.33 The tumor protein 53 (TP53) is one of three genes found to be frequently mutated in BC35,36 and plays a crucial role in conserving DNA stability, preventing cancer, and cell cycle arrest.37 In various biological conditions, such as DNA damage and cellular stresses, p53 can lead to different responses that similarly involve apoptosis, senescence, DNA repair, and cell cycle arrest.38,39
Another underlying mechanism of cancer development is histone modifications.40 KAT2B is one of the KAT family members and regulates acetylation and transcriptional processes throughout several biological procedures, including cancer.41 However, its regulatory roles in BC need to be investigated more.
This manuscript aims to explore the levels of miR-141, TP53, E2F3, CDK3, and KAT2B genes expression in BC tissues and highlight their association with clinicopathological features, such as metastasis and histologic grade. Moreover, the relationship of these 4 genes based on bioinformatic data is studied. The correlation between the RNA level of the mentioned genes is also investigated.
Materials and Methods
Bioinformatics analysis
MicroRNA target prediction involves identifying potential target genes of miR-141-3p. The most common miR-141-3p target genes, identified through 3P-seq tags and predicted by databases like miRDB, TargetScan, and miRWalk, are considered. Studies have explored the relationship between miR-141 target gene expression changes and breast cancer, revealing key genes like SOX4, BMI1, OCT4, zeb1-zeb2, CDKN2, CCNE1, CDKN1B, MDM2, E2F3, CDK3, KAT2B, and TP53. Bioinformatics research using KEGG and REACTOME databases, along with other studies, indicates that miR-141-3p target genes E2F3, CDK3, TP53, and KAT2B are involved in the cyclin-dependent kinase pathways, specifically the CDK4/6 signaling pathway related to cell cycle control complexes. These genes play crucial roles in breast cancer and treatment resistance. However, their combined impact in Triple-Negative Breast Cancer (TNBC) and their associations with miR-141, as well as their influence on histologic tumor grade and metastasis in untreated breast cancer tissues, remain unexplored.
Furthermore, the investigation utilized STRING, GENEMANIA, and their outcomes to identify relationships among genes in various functions, especially in physical interactions, accounting for 77.64% of the interactions.
A protein-protein interaction (PPI) network was constructed using databases such as STRING, GeneMANIA, and HIPPIE to assess the potential interactions between protein-coding genes with a medium confidence score of 0.4 and a filter score of 0.89. The interactions of the four genes were analyzed using these databases, and the networks were visualized using Cytoscape software v3.10.1, highlighting both functional and physical associations between proteins.
Patient selection and sample collection
Forty-six fresh breast tissue samples (23 normal and 23 tumor breast specimens) from female patients without reception of any surgical and antitumor therapy were provided by the Cancer Institute of Imam Khomeini Hospital (Tumor Bank, Tehran, Iran). Sample diagnosis was conducted between March 2018 and September 2019. Supplementary Pathological parameters for samples comprising of age, TNM stage, histologic grade, lymph node metastasis, tumor size, vascular and lymph invasion, and immune histochemical conditions of p53, ER, PR, and HER-2 receptors underwent examinations in the Pathology Laboratory of Imam Hospital, Tehran, Iran. All samples were stored at liquid nitrogen and then preserved at -80℃ to avoid RNA degradation. Tissue samples were gathered with the written consent of all patients. All of the patients were in stage 2 or 3 of progressive BC.
RNA isolation and cDNA synthesis
Total RNA extraction from frigid breast tumors and normal specimens was carried out after homogenizing with Trizol based on the manufacturer’s Tripure Isolation Reagent (Roche, cat. No. 93956620, USA) protocols. NanoDropOnec Spectrophotometer (Thermo Scientific, USA) was utilized to measure RNA purity and concentration. Complementary DNA (cDNA) synthesis from the extracted RNA was executed by a cDNA synthesis kit (PARSTOUS, Molecular Biology Kit, Iran). Total microRNA was extracted with the miRNA Extractor kit (BIOBASIC, Molecular Biology Kit, Lot:SK8811-M611ROX, USA) under the manufacturer’s instructions. Complementary DNA (cDNA) synthesis from the extracted microRNA for has-miR-141 was performed using a cDNA synthesis kit (Notarkib, Molecular Biology Kit, Iran) with stem-loop RT primers. Table S1 presents the sequences of the stem-loop RT primers.
Real-time-quantitative polymerase chain reaction (RT-qPCR)
Design of Specific forward and reverse primers was performed by means of oligo7, primer3, and Primer-BLAST for target genes and internal control. Table S1 presents the studied genes primers sequences.
RT-qPCR test was conducted by SYBR Green qPCR Master Mix (NAT Biotech Co, RealTiQ, Cat Number: NAT0011, Iran) and the specific mRNA PCR primers. RT-qPCR tests with a final volume of 10 µl were performed by ROTOR GENE Real-Time PCR Detection System under the following condition and the PCR temperature protocol at 95 ℃ for 10 min and then 40 cycles of 95 ℃, 59 ℃, and 72℃ for 15, 20, and 20 s, respectively, and melting temperature from 60 ℃ to 95 ℃. For all samples, the RT-qPCR test was performed duplicate, and the threshold cycle (Ct) was used to define mRNA expression. GAPDH gene was employed as an internal control gene to the mRNA expression normalization. The 2-ΔΔCt equation was utilized to estimate the relative expression level of the genes.
The microRNA expression analyses were conducted through an RT-qPCR test by SYBR Green qPCR Master Mix (NAT Biotech Co, RealTiQ, Cat Number: NAT0011, Iran) and the specific microRNA PCR primers. The specific forward primer was as follows: miR-141: 5’GCGCGTAACACTGTCTGGTA3’. The reverse primer was universal. RT-qPCR tests were conducted in ROTOR GENE Real-Time PCR Detection System with 10 µl final volume and under the following condition: 95 ℃ for10 min and then 40 cycles of 95 ℃ and 60 ℃ for 15 s and 60 s, respectively, and melting temperature from 60 ℃ to 95 ℃. For all samples, the RT-qPCR test was conducted duplicate, and to define miRNA expression, the threshold cycle (Ct) was used. Considering RNU6(U6) as the internal control expression level, the 2-ΔΔCt equation was employed to normalize the relative expression level.
Statistical analyses
IBM SPSS Statistics 26 (Armonk, NY, USA) was utilized to perform all statistical analyses. Normality was detemrined for each group using the Kolmogorov-Smirnov and Shapiro-Wilk normality test before any experiments were conducted. The Level of miR-141 expression was quantified by RT-qPCR in the breast tissue specimens, and RNU6 mRNA was used as the reference gene to normalize the results. Calculations of the fold changes were as the miRNA expression ratio to the internal control. Parametric and non-parametric tests were used (PAR: t-test, NPAR: Kruskal-Wallis test, Mann–Whitney U test) for comparisons of independent groups. The Pearson’s test was employed to determine if there is association amongst expression levels of these RNAs. P < 0.05 represented a statistically significant divergenece. The columnplots and scatterplots were made by GraphPad Prism 9 (GraphPad Software, La Jolla, CA).
Results
Bioinformatics analyses
As illustrated in Fig. 1, the String database showed a PPI network between all four candidate protein-coding genes, including 4 nodes and 5 edges highlighting P = 0.003 for the PPI enrichment. Genemania analysis demonstrated a PPI network between all four candidate genes as well. The resulting network is shown in Fig. 2. According to Genemania, 77.64% of interactions were physical, suggesting a relatively robust network between genes. Fig. 3 showes the PPI network of HIPPIE database regarding candidate genes. As it is demonstrated, four genes are in connection with each other and other protein coding genes. One interesting point is the multiple connection which was identified between TP53 and KAT2B in this data base.

Fig. 1.
The results of the interference of four studied genes are obtained from the STRING database and visualized using Cytoscape software v3.10.1. The blue circles in the legend represent nodes that are protein genes, while the lines represent the edges, which indicate interactions between proteins. The network comprises 4 nodes and 5 edges, with a PPI enrichment p-value of 0.002, indicating a strong interaction between candidate genes. The average node degree was 2.5.
.
The results of the interference of four studied genes are obtained from the STRING database and visualized using Cytoscape software v3.10.1. The blue circles in the legend represent nodes that are protein genes, while the lines represent the edges, which indicate interactions between proteins. The network comprises 4 nodes and 5 edges, with a PPI enrichment p-value of 0.002, indicating a strong interaction between candidate genes. The average node degree was 2.5.

Fig. 2.
The PPI network resulted from the Genemania database is visualized using Cytoscape software v3.10.1. The network displays highly interacting proteins with multiple interactions. The red and green circles represent candidate genes from the present study and their closely interacting proteins, respectively. PPI stands for Protein-Protein Interaction.
.
The PPI network resulted from the Genemania database is visualized using Cytoscape software v3.10.1. The network displays highly interacting proteins with multiple interactions. The red and green circles represent candidate genes from the present study and their closely interacting proteins, respectively. PPI stands for Protein-Protein Interaction.
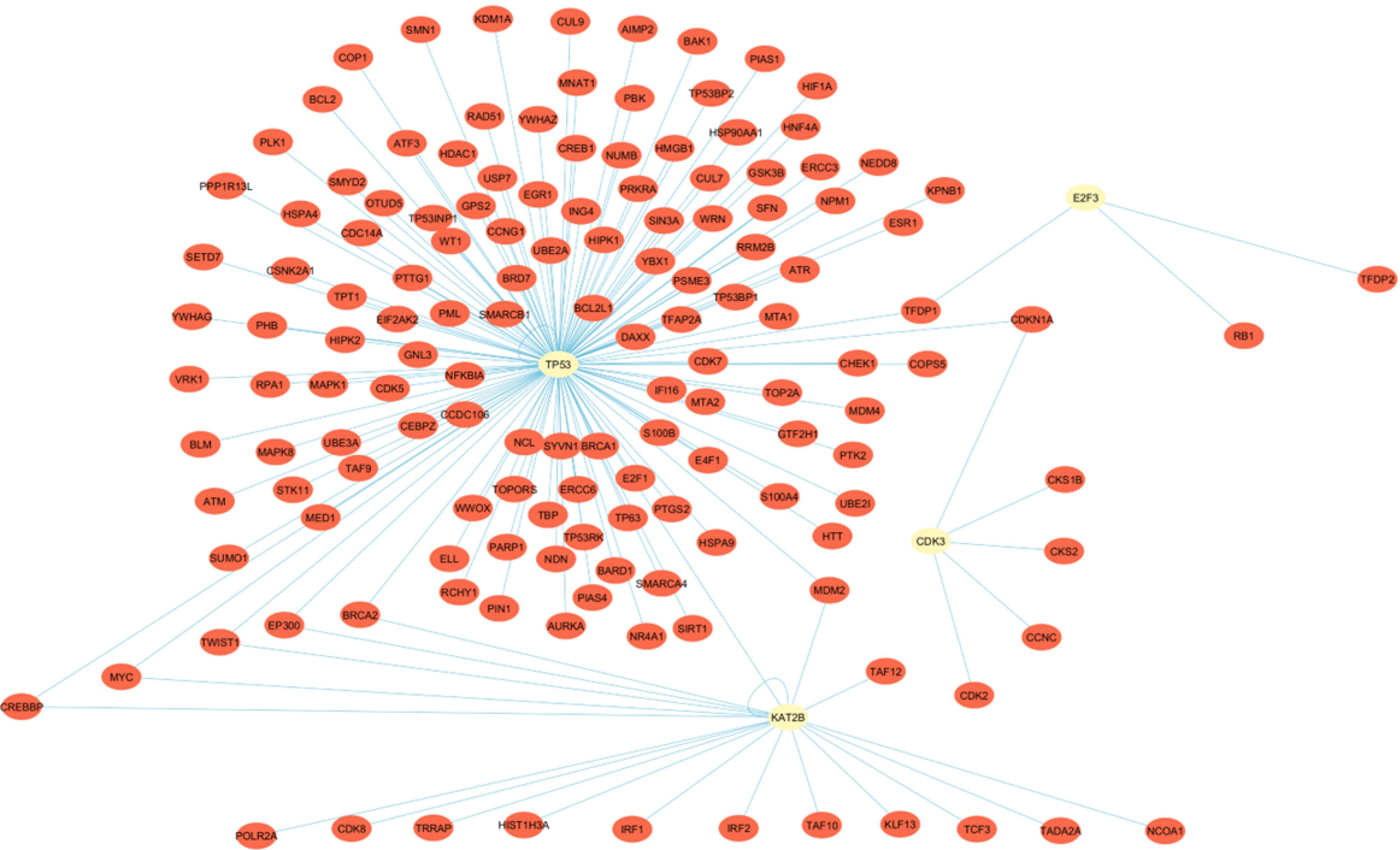
Fig. 3.
The PPI network of candidate genes was retrieved from the HIPPIE database with a score filter of 0.89 and visualized using Cytoscape software v3.10.1. As demonstrated, there are several interactions between candidate genes (yellow circles) with each other and other proteins (red circles). PPI stands for Protein-Protein Interaction.
.
The PPI network of candidate genes was retrieved from the HIPPIE database with a score filter of 0.89 and visualized using Cytoscape software v3.10.1. As demonstrated, there are several interactions between candidate genes (yellow circles) with each other and other proteins (red circles). PPI stands for Protein-Protein Interaction.
On the other hand, miRNA target prediction by means of miRDB, target scan, and miRWalk database demonstrated that E2F3 could be a potential target of miR-141-3p. Table S2 presents the list of predicted targets by Target scan for E2F3, showing the type of seed matching.
Overexpression of miR-141 in BC tissue samples
The present project examined the miR-141 differential expression by RT-qPCR in specimens of 23 BC patients in comparison to those of 23 normal breast tissue samples. The relative expression analyses were conducted on 23 normal breast tissue samples. Expression in different samples was normalized to U6, which is an appropriate housekeeping gene. Table S3 shows the clinical characteristics of the patients. The results demonstrated significant overexpression (fold change: 7.8, P = 0.0005) of miR-141 in BC tissues compared with normal ones (Fig. 4).

Fig. 4.
The expression levels of miR-141 in tumor and normal breast tissue samples were evaluated. Baseline levels of miR-141 in tumor and normal breast tissues were determined using real-time PCR. Statistical analysis was performed using the Mann–Whitney U test. ***P ≤ 0.001.
.
The expression levels of miR-141 in tumor and normal breast tissue samples were evaluated. Baseline levels of miR-141 in tumor and normal breast tissues were determined using real-time PCR. Statistical analysis was performed using the Mann–Whitney U test. ***P ≤ 0.001.
Patients’ characteristics
Two out of 23 patients (8.69%) had well-differentiated cancer tissues (Grade I), while there were 10 cases (43.47%) with moderately differentiated (Grade II) and 9 cases (39.13%) with poorly differentiated cancer tissues (Grade III). More information about clinicopathological characteristics is presented in (Supplementary file 1, Table S3).
Expression pattern of E2F3, CDK3, TP53, and KAT2B in BC tissues compared to normal tissues
TP53, E2F3, CDK3, and KAT2B differential expression was investigated using RT-qPCR in 23 BC tissues in comparison to those of 23 normal tissues. Our findings demonstrated the significant overexpression of three candidate genes of this study (E2F3, CDK3, and KAT2B) in BC compared with normal tissues (Fig. 5). Table 1 presents the Fold change of each gene and its respective P value.
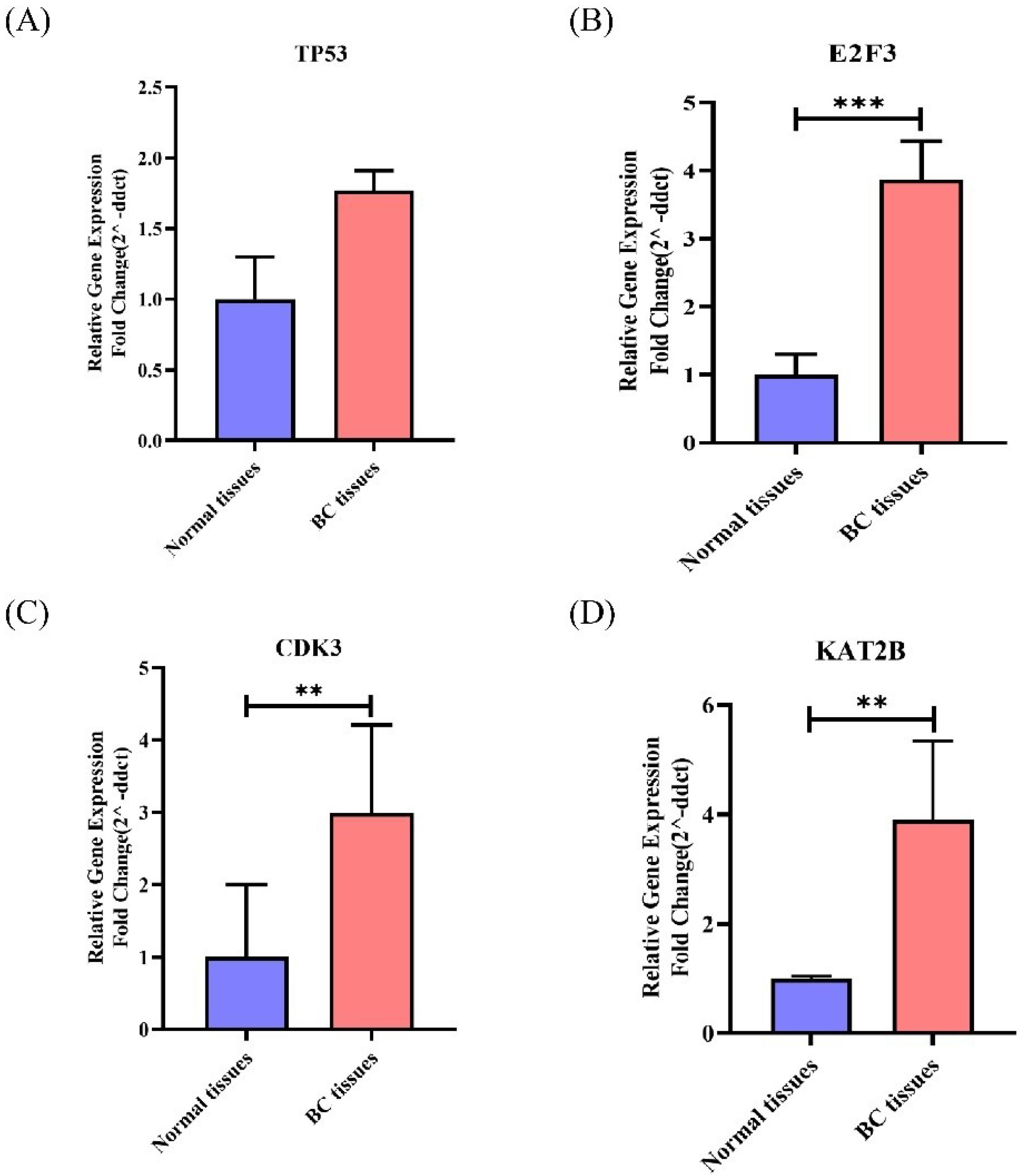
Fig. 5.
Expression levels of TP53, E2F3, CDK3, KAT2B in BC tissues. Higher expression of TP53, E2F3, CDK3, and KAT2B was observed in BC tissues than in benign breast specimens, which all were statistically significant (except TP53).TP53(A), E2F3(B), CDK3(C), KAT2B(D). **P ≤ 0.01 and ***P ≤ 0.001.
.
Expression levels of TP53, E2F3, CDK3, KAT2B in BC tissues. Higher expression of TP53, E2F3, CDK3, and KAT2B was observed in BC tissues than in benign breast specimens, which all were statistically significant (except TP53).TP53(A), E2F3(B), CDK3(C), KAT2B(D). **P ≤ 0.01 and ***P ≤ 0.001.
Table 1.
Mean expression of genes in tumor tissues compared to normal breast tissues
|
Genes
|
Normal Mean
2-ΔΔCt |
Tumor Mean
2-ΔΔCt |
Pattern
|
P
value
|
| TP53 |
1.3 |
1.7 |
Upregulated |
0. 2 |
| E2F3 |
1.2 |
3.8 |
Upregulated |
0.0008 |
| CDK3 |
1.3 |
2.9 |
Upregulated |
0.008 |
| KAT2B |
0.1 |
3.9 |
Upregulated |
0.03 |
| miR-141 |
1.7 |
7.8 |
Upregulated |
0.0005 |
The expression pattern of miR-141 and selected genes in BC tissues and their association with clinicopathological characteristics
Additional analyses were conducted to determine whether there was any relationship among the miR-141 mean expression and selected genes with the clinical and pathological characteristics of BC patients. The expression levels of miR-141, E2F3, CDK3, and KAT2B were detected to be significantly related to the histologic grading of the BC tumor (Table 2), highlihgting significantly greater expression levels of miR-141 in Grade 3 (P = 0.002) than in Grade 2 (P = 0.003) of BC patients. Similarly, the expression levels of E2F3 were significantly higher in Grade 3 (P = 0.001) than in Grade 2 (P = 0.008) BC patients. However, KAT2B and CDK3 had significantly higher expression levels in Grade 3 tissue samples (P = 0.03 and P = 0.01, respectively) but did not reveal a significant divergence in Grade 2. In other words, all the studied genes except TP53 were differentially expressed in Grade 3 specimens.
Table 2.
Relationship between RNA expression level of miR-141 and target genes withclinicopathological features from BC tissues
|
Clinicopathological characteristics
|
Relevant expression of genes (2-ΔΔCt), P value |
|
TP53
|
E2F3
|
CDK3
|
KAT2B
|
miR-141
|
| Histologic grade (Tumor differentiation) |
|
| Grade I (Low) |
0.6 |
0.4 |
0.02*
|
|
0.6 |
| Grade II (Moderately) |
0.2 |
0.008* |
0.1 |
0.2 |
0.003* |
| Grade III (poorly) |
0.6 |
0.001* |
0.01* |
0.03* |
0.002* |
| HER2 status |
|
| Positive |
0.6 |
0.001* |
0.3 |
0.6 |
0.4 |
| Negative |
0.1 |
0.02* |
0.03* |
0.09 |
0.7 |
| Metastatic |
|
| M0 (Regional) |
0.2 |
0.002* |
0.01* |
0.008* |
0.000* |
| M1 (Distant) |
|
0.03* |
|
0.07 |
|
* Statistically significant (P < 0.05).
Furthermore, E2F3 and CDK3 expression levels were significantly related to HER2 status in BC tissues. The mean expression levels of E2F3 were significantly higher in BC patients having HER2 + than those with HER2-. The Mean expression levels of CDK3 were significantly related to HER2- (absence of HER2).
Also, E2F3, CDK3, KAT2B, and miR-141 expression levels were significantly related to M0 metastasis status (i.e., no distant cancer spread was found). The mean expression level of E2F3 was considerably greater in BC patients having M0 status than those with M1 status.
Several significant correlations were observed between the RNA expression levels of the selected genes in BC (Table 3) and adjacent normal tissues (Table 4), as demonstrated by Pearson correlation values.
Table 3.
Correlation coefficients according to pearson between RNA expression of genes for all adjacent normal tissues from BC patients
|
|
E2F3
|
CDK3
|
KAT2B
|
TP53
|
| MiR-141 |
0.51 |
0.99* |
|
0.95** |
| E2F3 |
1 |
0.92 |
-0.52 |
0.97* |
| CDK3 |
|
1 |
|
0.93 |
| KAT2B |
|
|
1 |
|
** Correlation is significant at the 0.01 level.* Correlation is significant at the 0.05 level.
Table 4.
Correlation coefficients according to pearson between RNA expression of genes for all tumor tissues from BC patients
|
|
E2F3
|
CDK3
|
KAT2B
|
TP53
|
| MiR-141 |
0.07 |
-0.78* |
0.36 |
0.2 |
| E2F3 |
1 |
-0.12 |
-0.96** |
0.31 |
| CDK3 |
|
1 |
0.51 |
0.70 |
| KAT2B |
|
|
1 |
0.005 |
** Correlation is significant at the 0.01 level.
* Correlation is significant at the 0.05 level.
A negative correlation was reported between miR-141 and CDK3 mRNA expression levels in BC tissues (P = 0.03, rs: -0.78) (Fig. 6A). Evidence also revealed a positive correlation between miR-141 and CDK3 expression levels in normal tissues (P = 0.03, rs:0.99) (Fig. 6A).
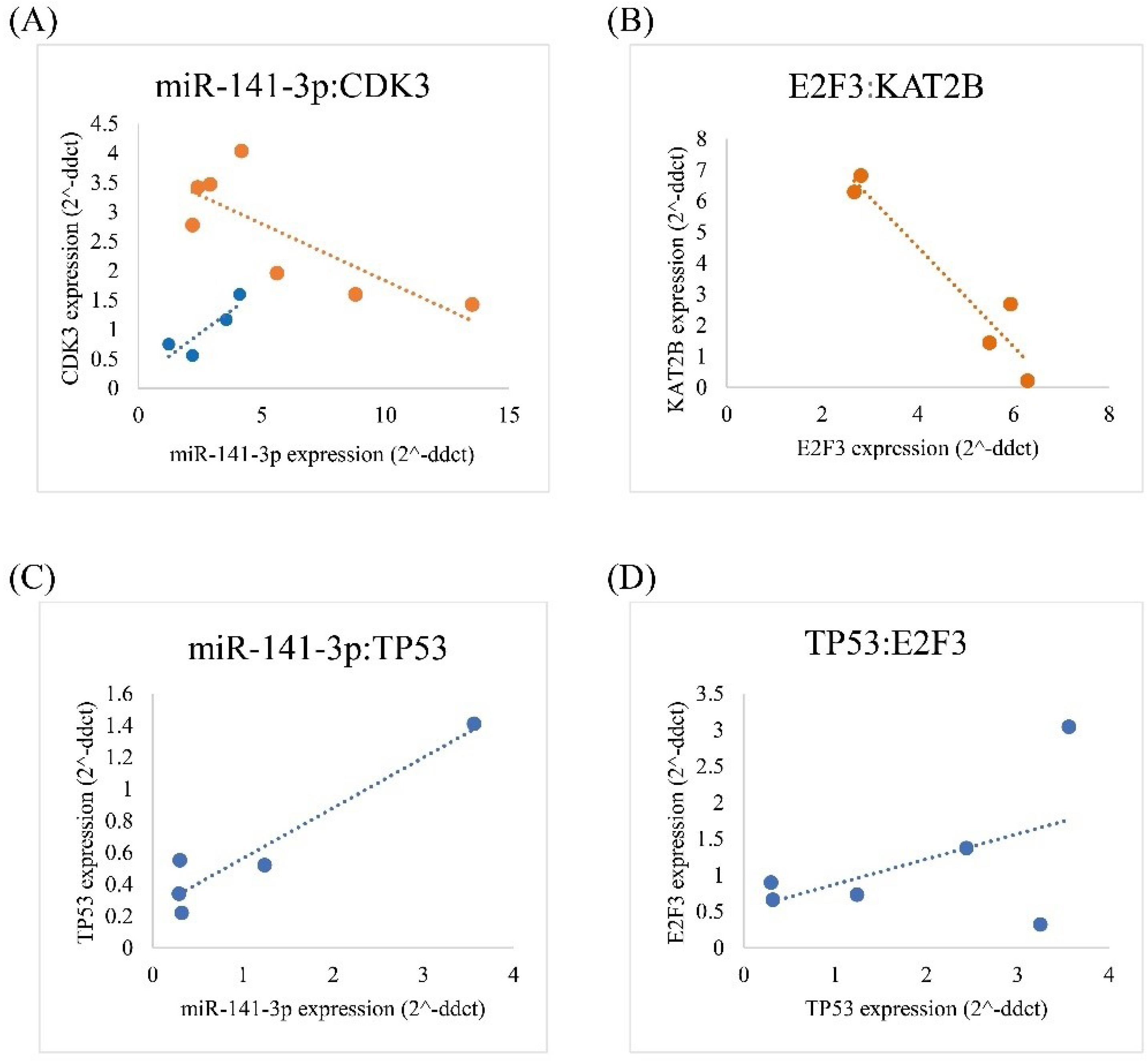
Fig. 6.
Significant correlations for expression levels between CDK3 and KAT2B with ncRNA (miR-141-3p). Correlations between CDK3 and miR-141-3p (A), KAT2B and E2F3 (B), TP53 and miR-141-3p (C), E2F3 and TP53 (D). (The red color indicate tumor tissues, while the blue color indicate normal tissues).
.
Significant correlations for expression levels between CDK3 and KAT2B with ncRNA (miR-141-3p). Correlations between CDK3 and miR-141-3p (A), KAT2B and E2F3 (B), TP53 and miR-141-3p (C), E2F3 and TP53 (D). (The red color indicate tumor tissues, while the blue color indicate normal tissues).
The expression levels of E2F3 mRNA and KAT2B had negative correlations in BC (P = 0.01, rs: -0.96) (Fig. 6B).
In addition, The miR-141 expression values were positively correlated with the TP53 expression values in adjacent normal tissues (P = 0.01, rs: 0.95) (Fig. 6C). Moreover the expression values of E2F3 were positively correlated with the expression values of TP53 in adjacent normal tissues (P = 0.02, rs: 0.97) (Fig. 6D).
Discussion
Most of the miRNAs in cancer research have been categorized based on their procedure involvement, i.e., oncogenic, apoptotic, and metastatic. The miR-200 family members (including miR-141) are usually categorized as metastasis, invasion, and treatment responses.42 Different types of cancer are regulated by intracellular miRNAs that act as underlying mechanisems of controling caretaker and gatekeeper genes. 7 However, fewer studies of miR-200 family members have been attributed to miR-141. Noteworthy, some good research has documented the importance of miR-141 in BC, as pointed out by Debeb et al and Taha et al, indicating a close association between the high expression levels of miR-181b1-5p and miR-141-3p with aggressive BC.43,44 Various expressions of miR-141 as an epigenetic alteration have a regulatory role in different steps of cancer such as metastasis, invasion, recurrence, and resistance to drugs.16 This study disclosed increased expression levels of miR-141 in the BC compared to normal patients, associated with regional metastasis. Similarly, Debeb et al. reported that miR-141 was upregulated in BC tissues.43 Other studies also documented the overexpression of all miR-200 family members, including miR-141, in breast tumors vs. normal breast tissues.45 The other point, requiring discussion here, was the finding that overexpression of miR-141 suppressed distant metastasis in all investigated specimens. The present outcome aligns with some prior studies indicating that downregulation of serum miR200c/141 was associated with tumor metastasis in BC.20 However, there are conflicting reports on miR-141 expression in BC tumor tissues, as Li et al reported downregulation of miR-141 in breast tumor tissue.46
Our results indicated that the E2F3 mRNA expression was incremented in human BC compared to normal tissues and was more significant in grade III samples. Consistent with our result, other studies reported the overexpression of E2F3 in BC.47 According to Shokrollahzade et al, the expression of E2F3 as the target of miR-141 increased in breast cancer cell lines, but this overexpression was not significant.7 In addition, Chen et al showed that the Brachyury/E2F3 axis might be a diagnostic biomarker, and Knockdown of E2F3 diminished BC cell growth and migration.48 Moreover, according to Jusino et al, the expression of SGO1 and E2F3 increased in invasive ductal BC compared with normal breast tissues, which correlated significantly with poor survival outcomes, epithelial-to-mesenchymal transition, and metastasis.32 Furthermore, we reported the protein-protein interaction between E2F3 and P53, and the positive correlation between them. These data are in good agreement with previous studies that indicate an association between E2F3 and P53. Reports have confirmed that E2F3 plays some regulatory roles in ARF/p53 pathway and p53-p21CIP1 Axis.49,50 Also, our results demonstrated a protein-protein interaction and a negative correlation between E2F3 and KAT2B, which is in line with the results obtained by Pio et al, reporting the E2F3 and KAT2B in one cluster as targets of several microRNAs.51 Targeting E2F3 by miR-141 has been reported in other conditions as well. Xue et al showed that miR-141 inhibited the osteogenic differentiation of bone marrow-derived mesenchymal stem cells in osteonecrosis of the femoral head by targeting E2F3.52 Besides, another study, conducted by Zhou et al, illustrated that gastric cancer was inhibited by the interaction of miR-141 with long noncoding RNA MEG3 that down-regulated E2F3 expression.53
Another investigated gene in this study was CDK3, which was upregulated in tumor compared to normal breast tissues. In agreement with our result, Zhang et al demonstrated that CDK3 was overexpressed in breast cancer cell lines, comprising of MDA-MB-231 And MCF7 cell lines.54 Cao et al showed that the overexpression of CDK3 as a target of miR-4469 decreased metastasis, invasion, and migration by impeding the Wnt/β-catenin signaling pathway.33 It was found that CDK3 and miR-141 were negatively correlated in tumor tissues but positively correlated in normal tissues. This is the first study that reports the correlations between CDK3 and miR-141, to the best of our knowledge. In addition, the investigated dysregulation of E2F3, CDK3, and TP53, which are important genes in the cell cycle, emphasizes the potential role of cell cycle malfunction during BC development or metastasis.
As reported in this study, miR-141 is upregulated in breast tumor tissues. Moreover, the results showed the overexpression of KAT2B, which is an acetyl-transferase p300/CBP-associated factor. In agreement with our findings, there an investigation reported the assocciation of miR-141 and KAT2B. According to Zhang et al, higher circulating miR-141 and miR-200c levels are potential biomarkers to detect breast cancer metastasis in early stages. They also demonstrated miR-200c/141 expression regulation through FOXP3-KAT2B proteins, which bind to the miR-200c/141 promoter and induce miR-200c/141 overexpression.20 Additionally, our study reported a protein-protein interaction between TP53 and KAT2B. In agreement with our study, Elengoe et al showed an interaction network based on the TP53 protein and identified genes involved in the tumor development process. One of the identified genes was KAT2B with molecular functions, such as enzyme binding role and transcription factor binding role.55
Our findings also demonstrated overexpression of TP53, but the difference between tumor and normal breast tissues was not significant. However, this result is in line with a cohort study of 278 stage I-III triple-negative breast invasive ductal carcinoma patients, demonstrating associations between TP53 overexpression in 58.6% (163/278) of patients which was associated with worse overall survival (Log-rank test P = 0.025).56 In this study, we found that TP53 and miR-141 significantly correlated in normal tissues. In the same vein, Zhou et al demonstrated a significant correlation between miR-141-3p and p53 expression in gliomas compared with normal brain tissue.57 Another study indicated the regulatory role of P53 on miR-141 in hepatocellular carcinomas. It is reported that miR-141 expression level was high in cell lines with wild type of P53 gene, but the expression level of miR-141 decreased significantly in mutant p53 cell lines, suggesting the regulatory effects between P53 and miR-141.58
Conclusion
The findings of this study documented overexpression of miR-141, E2F3, CDK3, TP53, and KAT2B in BC tissues. It was also demonstrated that overexpression of some of these genes, was associated with a higher grade of the disease and its regional metastasis. Thus, current data highlighted indicators that could be a useful marker of further disease progression and clinicopathologically relevant to BC patients. Moreover, the predicted interactions and correlations between the investigated genes in this study provide new information of the possible network of these genes in BC and might imply regulatory effects. The results of this study have the potential to be applied for the assessment of higher disease levels in BC patients, contributing as a useful criteria for clinicians to determine more appropriate interventions. Therefore, these promissing data warrant more investigations on miR-141 and related genes to fully underestand their role in BC.
Research Highlights
What is the current knowledge?
-
BC is the second reason of mortality by cancer in females globally.
-
Many human malignancies, such as breast cancer, express miR-141 as one of the members of the miR-200 family.
-
Key genes are engaged in the cell cycle that is differentially expressed during cancer development or metastasis that TP53, E2F3, CDK3, and KAT2B are examples of these identified genes.
What is new here?
-
Overexpression of miR-141, E2F3, CDK3, and KAT2B was correlated with aggressive breast cancer, suggesting regulatory effects of these genes in breast cancer.
-
Overexpression of some of the studied genes was associated with two pathological features, including a higher grade of the disease and its regional metastasis.
-
A significant correlation was detected between RNA level of miR-141 with CDK3 and E2F3 with KAT2B in this study.
Acknowledgments
This work was elicited from the MSc. thesis of Sepideh Ebrahimian Vargahan that was supported by Project number (18320) from Iran University of Medical Science Research Affairs. Our sincere thanks go out to the support provided by the Department of Medical Biotechnology, school of Allied Medical Sciences, Iran University of Medical Sciences, Tehran, Iran.
Competing Interests
The authors claim no conflict of interests.
Ethical Statement
The current project obtained the approval of the Ethics Committee of Iran University of Medical Sciences. Consent forms were filled out by all patients before using their tissue samples based on the Declaration of Helsinki (DoH).
Supplementary files
Supplementary file 1 contains Tables S1-S3.
(pdf)
References
- Turashvili G, Brogi E. Tumor heterogeneity in breast cancer. Front Med (Lausanne) 2017; 4:227. doi: 10.3389/fmed.2017.00227 [Crossref] [ Google Scholar]
- Afzal M, Ameeduzzafar Ameeduzzafar, Alharbi KS, Alruwaili NK, Al-Abassi FA, Al-Malki AAL. Nanomedicine in treatment of breast cancer - a challenge to conventional therapy. Semin Cancer Biol 2021; 69:279-92. doi: 10.1016/j.semcancer.2019.12.016 [Crossref] [ Google Scholar]
- Choy TK, Wang CY, Phan NN, Khoa Ta HD, Anuraga G, Liu YH. Identification of dipeptidyl peptidase (DPP) family genes in clinical breast cancer patients via an integrated bioinformatics approach. Diagnostics (Basel) 2021; 11:1204. doi: 10.3390/diagnostics11071204 [Crossref] [ Google Scholar]
- Li XX, Wang LJ, Hou J, Liu HY, Wang R, Wang C. Identification of long noncoding RNAs as predictors of survival in triple-negative breast cancer based on network analysis. Biomed Res Int 2020; 2020:8970340. doi: 10.1155/2020/8970340 [Crossref] [ Google Scholar]
- Wang L, Zhang S, Wang X. The metabolic mechanisms of breast cancer metastasis. Front Oncol 2020; 10:602416. doi: 10.3389/fonc.2020.602416 [Crossref] [ Google Scholar]
- Eini M, Parsi S, Barati M, Bahramali G, Alizadeh Zarei M, Kiani J. Bioinformatic investigation of microRNA-802 target genes, protein networks, and its potential prognostic value in breast cancer. Avicenna J Med Biotechnol 2022; 14:154-64. doi: 10.18502/ajmb.v14i2.8882 [Crossref] [ Google Scholar]
- Shokrollahzade S, Hosseini A, Safa M, Sarhadi S. Differential expression of EGFR, MAP2K4 and E2F3 genes as targets of miR-141 and its association with immune system pathway. J Cell Mol Res 2017; 9:6-15. doi: 10.22067/jcmr.v0i0.58965 [Crossref] [ Google Scholar]
- Liu W, Ma C, Xu H, Wang L, Xu W, Zhang H. miR-184-5p inhibits cell proliferation, invasion and predicts prognosis of clear cell renal cell carcinoma by targeting NUS1 dehydrodolichyl diphosphate synthase subunit: results from large-scale comprehensive identification and validation. J Cancer 2022; 13:1398-409. doi: 10.7150/jca.58053 [Crossref] [ Google Scholar]
- Jo H, Shim K, Jeoung D. Potential of the miR-200 family as a target for developing anti-cancer therapeutics. Int J Mol Sci 2022; 23:5881. doi: 10.3390/ijms23115881 [Crossref] [ Google Scholar]
- Davey MG, Davies M, Lowery AJ, Miller N, Kerin MJ. The role of microRNA as clinical biomarkers for breast cancer surgery and treatment. Int J Mol Sci 2021; 22:8290. doi: 10.3390/ijms22158290 [Crossref] [ Google Scholar]
- Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet 2009; 10:704-14. doi: 10.1038/nrg2634 [Crossref] [ Google Scholar]
- Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D. MicroRNA expression profiles classify human cancers. Nature 2005; 435:834-8. doi: 10.1038/nature03702 [Crossref] [ Google Scholar]
- Choi SK, Kim HS, Jin T, Hwang EH, Jung M, Moon WK. Overexpression of the miR-141/200c cluster promotes the migratory and invasive ability of triple-negative breast cancer cells through the activation of the FAK and PI3K/AKT signaling pathways by secreting VEGF-A. BMC Cancer 2016; 16:570. doi: 10.1186/s12885-016-2620-7 [Crossref] [ Google Scholar]
- Cavallari I, Ciccarese F, Sharova E, Urso L, Raimondi V, Silic-Benussi M. The miR-200 family of microRNAs: fine tuners of epithelial-mesenchymal transition and circulating cancer biomarkers. Cancers (Basel) 2021; 13:5874. doi: 10.3390/cancers13235874 [Crossref] [ Google Scholar]
- Shi L, Sun HJ, Zeng JJ, Liang ZQ, Lin YH, Huang SN. Evaluation of miR-141-3p over-expression in ovarian cancer. Electron J Biotechnol 2022; 58:14-24. doi: 10.1016/j.ejbt.2022.04.006 [Crossref] [ Google Scholar]
- Gao Y, Feng B, Han S, Zhang K, Chen J, Li C. The roles of microRNA-141 in human cancers: from diagnosis to treatment. Cell Physiol Biochem 2016; 38:427-48. doi: 10.1159/000438641 [Crossref] [ Google Scholar]
- Yang G, Lu Z, Meng F, Wan Y, Zhang L, Xu Q. Circulating miR-141 as a potential biomarker for diagnosis, prognosis and therapeutic targets in gallbladder cancer. Sci Rep 2022; 12:10072. doi: 10.1038/s41598-022-13430-8 [Crossref] [ Google Scholar]
- Liu C, Li Z, Ni L. Analysis of the significance of miR-141 and CD147 expression in bladder cancer cells and its relationship with tumor grade. Contrast Media Mol Imaging 2022; 2022:1335441. doi: 10.1155/2022/1335441 [Crossref] [ Google Scholar]
- Simpson KE, Watson KL, Moorehead RA. Elevated expression of miR-200c/141 in MDA-MB-231 cells suppresses MXRA8 levels and impairs breast cancer growth and metastasis in vivo. Genes (Basel) 2022; 13:691. doi: 10.3390/genes13040691 [Crossref] [ Google Scholar]
- Zhang G, Zhang W, Li B, Stringer-Reasor E, Chu C, Sun L. MicroRNA-200c and microRNA- 141 are regulated by a FOXP3-KAT2B axis and associated with tumor metastasis in breast cancer. Breast Cancer Res 2017; 19:73. doi: 10.1186/s13058-017-0858-x [Crossref] [ Google Scholar]
- Li J, Huang G, Ren C, Wang N, Sui S, Zhao Z. Identification of differentially expressed genes-related prognostic risk model for survival prediction in breast carcinoma patients. Aging (Albany NY) 2021; 13:16577-99. doi: 10.18632/aging.203178 [Crossref] [ Google Scholar]
- Hsu JB, Lee TY, Cheng SJ, Lee GA, Chen YC, Le NQK. Identification of differentially expressed genes in different glioblastoma regions and their association with cancer stem cell development and temozolomide response. J Pers Med 2021; 11:1047. doi: 10.3390/jpm11111047 [Crossref] [ Google Scholar]
- Ge Y, Zhang C, Xiao S, Liang L, Liao S, Xiang Y. Identification of differentially expressed genes in cervical cancer by bioinformatics analysis. Oncol Lett 2018; 16:2549-58. doi: 10.3892/ol.2018.8953 [Crossref] [ Google Scholar]
- Annapurna SD, Pasumarthi D, Pasha A, Doneti R, Sheela B, Botlagunta M. Identification of differentially expressed genes in cervical cancer patients by comparative transcriptome analysis. Biomed Res Int 2021; 2021:8810074. doi: 10.1155/2021/8810074 [Crossref] [ Google Scholar]
- Bai B, Chen Q, Jing R, He X, Wang H, Ban Y, et al. Molecular basis of prostate cancer and natural products as potential chemotherapeutic and chemopreventive agents. Front Pharmacol 2021. 12: 738235. 10.3389/fphar.2021.738235.
- Liu J, Peng Y, Wei W. Cell cycle on the crossroad of tumorigenesis and cancer therapy. Trends Cell Biol 2022; 32:30-44. doi: 10.1016/j.tcb.2021.07.001 [Crossref] [ Google Scholar]
- Kontomanolis EN, Koutras A, Syllaios A, Schizas D, Kalagasidou S, Pagkalos A. Basic principles of molecular biology of cancer cell-molecular cancer indicators. J BUON 2021; 26:1723-34. [ Google Scholar]
- DeGregori J. The genetics of the E2F family of transcription factors: shared functions and unique roles. Biochim Biophys Acta 2002; 1602:131-50. doi: 10.1016/s0304-419x(02)00051-3 [Crossref] [ Google Scholar]
- Gao Y, Feng B, Lu L, Han S, Chu X, Chen L. MiRNAs and E2F3: a complex network of reciprocal regulations in human cancers. Oncotarget 2017; 8:60624-39. doi: 10.18632/oncotarget.17364 [Crossref] [ Google Scholar]
- Leone G, DeGregori J, Yan Z, Jakoi L, Ishida S, Williams RS. E2F3 activity is regulated during the cell cycle and is required for the induction of S phase. Genes Dev 1998; 12:2120-30. doi: 10.1101/gad.12.14.2120 [Crossref] [ Google Scholar]
- Wu L, Wan S, Li J, Xu Y, Lou X, Sun M. Expression and prognostic value of E2F3 transcription factor in non-small cell lung cancer. Oncol Lett 2021; 21:411. doi: 10.3892/ol.2021.12672 [Crossref] [ Google Scholar]
- Jusino S, Rivera-Rivera Y, Chardón-Colón C, Ruiz-Justiz AJ, Vélez-Velázquez J, Isidro A. E2F3 drives the epithelial-to-mesenchymal transition, cell invasion, and metastasis in breast cancer. Exp Biol Med (Maywood) 2021; 246:2057-71. doi: 10.1177/15353702211035693 [Crossref] [ Google Scholar]
- Cao T, Xiao T, Huang G, Xu Y, Zhu JJ, Wang K. CDK3, target of miR-4469, suppresses breast cancer metastasis via inhibiting Wnt/β-catenin pathway. Oncotarget 2017; 8:84917-27. doi: 10.18632/oncotarget.18171 [Crossref] [ Google Scholar]
- Ghafouri-Fard S, Khoshbakht T, Hussen BM, Dong P, Gassler N, Taheri M. A review on the role of cyclin dependent kinases in cancers. Cancer Cell Int 2022; 22:325. doi: 10.1186/s12935-022-02747-z [Crossref] [ Google Scholar]
- Hainaut P, Hollstein M. p53 and human cancer: the first ten thousand mutations. Adv Cancer Res 2000; 77:81-137. doi: 10.1016/s0065-230x(08)60785-x [Crossref] [ Google Scholar]
- Kim JY, Park K, Jung HH, Lee E, Cho EY, Lee KH. Association between mutation and expression of TP53 as a potential prognostic marker of triple-negative breast cancer. Cancer Res Treat 2016; 48:1338-50. doi: 10.4143/crt.2015.430 [Crossref] [ Google Scholar]
- Berke TP, Slight SH, Hyder SM. Role of reactivating mutant p53 protein in suppressing growth and metastasis of triple-negative breast cancer. Onco Targets Ther 2022; 15:23-30. doi: 10.2147/ott.s342292 [Crossref] [ Google Scholar]
- Koo N, Sharma AK, Narayan S. Therapeutics targeting p53-MDM2 interaction to induce cancer cell death. Int J Mol Sci 2022; 23:5005. doi: 10.3390/ijms23095005 [Crossref] [ Google Scholar]
- Chen J. The cell-cycle arrest and apoptotic functions of p53 in tumor initiation and progression. Cold Spring Harb Perspect Med 2016; 6:a026104. doi: 10.1101/cshperspect.a026104 [Crossref] [ Google Scholar]
- Yang Y, Zhang M, Wang Y. The roles of histone modifications in tumorigenesis and associated inhibitors in cancer therapy. J Natl Cancer Cent 2022; 2:277-90. doi: 10.1016/j.jncc.2022.09.002 [Crossref] [ Google Scholar]
- Li L, Zhang J, Cao S. Lysine acetyltransferase 2B predicts favorable prognosis and functions as anti-oncogene in cervical carcinoma. Bioengineered 2021; 12:2563-75. doi: 10.1080/21655979.2021.1935525 [Crossref] [ Google Scholar]
- Taylor MA, Schiemann WP. Therapeutic opportunities for targeting microRNAs in cancer. Mol Cell Ther 2014; 2:1-13. doi: 10.1186/2052-8426-2-30 [Crossref] [ Google Scholar]
- Taha M, Mitwally N, Soliman AS, Yousef E. Potential diagnostic and prognostic utility of miR-141, miR-181b1, and miR-23b in breast cancer. Int J Mol Sci 2020; 21:8589. doi: 10.3390/ijms21228589 [Crossref] [ Google Scholar]
- Debeb BG, Lacerda L, Anfossi S, Diagaradjane P, Chu K, Bambhroliya A. miR-141-mediated regulation of brain metastasis from breast cancer. J Natl Cancer Inst 2016; 108:djw026. doi: 10.1093/jnci/djw026 [Crossref] [ Google Scholar]
- Fontana A, Barbano R, Dama E, Pasculli B, Rendina M, Morritti MG. Combined analysis of miR-200 family and its significance for breast cancer. Sci Rep 2021; 11:2980. doi: 10.1038/s41598-021-82286-1 [Crossref] [ Google Scholar]
- Li P, Xu T, Zhou X, Liao L, Pang G, Luo W. Downregulation of miRNA-141 in breast cancer cells is associated with cell migration and invasion: involvement of ANP32E targeting. Cancer Med 2017; 6:662-72. doi: 10.1002/cam4.1024 [Crossref] [ Google Scholar]
- Vimala K, Sundarraj S, Sujitha MV, Kannan S. Curtailing overexpression of E2F3 in breast cancer using siRNA (E2F3)-based gene silencing. Arch Med Res 2012; 43:415-22. doi: 10.1016/j.arcmed.2012.08.009 [Crossref] [ Google Scholar]
- Chen M, Liu J, Liang X, Huang Y, Yang Z, Lu P. Knockdown of brachyury suppresses breast cancer cell proliferation and migration via targeting E2F3. J Oncol 2022; 2022:7913067. doi: 10.1155/2022/7913067 [Crossref] [ Google Scholar]
- Ginsberg D. E2F3-a novel repressor of the ARF/p53 pathway. Dev Cell 2004; 6:742-3. doi: 10.1016/j.devcel.2004.05.012 [Crossref] [ Google Scholar]
- Sharma N, Timmers C, Trikha P, Saavedra HI, Obery A, Leone G. Control of the p53-p21CIP1 Axis by E2f1, E2f2, and E2f3 is essential for G1/S progression and cellular transformation. J Biol Chem 2006; 281:36124-31. doi: 10.1074/jbc.M604152200 [Crossref] [ Google Scholar]
- Pio G, Ceci M, D'Elia D, Loglisci C, Malerba D. A novel biclustering algorithm for the discovery of meaningful biological correlations between microRNAs and their target genes. BMC Bioinformatics 2013; 14 Suppl 7:S8. doi: 10.1186/1471-2105-14-s7-s8 [Crossref] [ Google Scholar]
- Xue F, Wu J, Feng W, Hao T, Liu Y, Wang W. MicroRNA-141 inhibits the differentiation of bone marrow-derived mesenchymal stem cells in steroid-induced osteonecrosis via E2F3. Mol Med Rep 2022; 26:234. doi: 10.3892/mmr.2022.12750 [Crossref] [ Google Scholar]
- Zhou X, Ji G, Ke X, Gu H, Jin W, Zhang G. MiR-141 inhibits gastric cancer proliferation by interacting with long noncoding RNA MEG3 and down-regulating E2F3 expression. Dig Dis Sci 2015; 60:3271-82. doi: 10.1007/s10620-015-3782-x [Crossref] [ Google Scholar]
- Zhang Z, Huang A, Zhang A, Zhou C. HuR promotes breast cancer cell proliferation and survival via binding to CDK3 mRNA. Biomed Pharmacother 2017; 91:788-95. doi: 10.1016/j.biopha.2017.04.063 [Crossref] [ Google Scholar]
- Elengoe A, Hamdan S. Mapping of TP53 protein network using cytoscape software. Pharm Biomed Res 2018; 4:9-25. doi: 10.18502/pbr.v4i3.539 [Crossref] [ Google Scholar]
- Li JP, Zhang XM, Zhang Z, Zheng LH, Jindal S, Liu YJ. Association of p53 expression with poor prognosis in patients with triple-negative breast invasive ductal carcinoma. Medicine (Baltimore) 2019; 98:e15449. doi: 10.1097/md.0000000000015449 [Crossref] [ Google Scholar]
- Zhou X, Wu W, Zeng A, Nie E, Jin X, Yu T. MicroRNA-141-3p promotes glioma cell growth and temozolomide resistance by directly targeting p53. Oncotarget 2017; 8:71080-94. doi: 10.18632/oncotarget.20528 [Crossref] [ Google Scholar]
- Kim T, Veronese A, Pichiorri F, Lee TJ, Jeon YJ, Volinia S. p53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J Exp Med 2011; 208:875-83. doi: 10.1084/jem.20110235 [Crossref] [ Google Scholar]